- About us
- Research
- Students & Teaching
- Seminars & Events
- Directories
- Booking Rooms & Equipment
- עברית
Home » Genes inherited from early ancestors are scattered among our chromosomes, more recent genes are often bunched close together
As evolution proceeded along the path from the first unicellular organisms through to us
humans, new life styles emerged. For a very long time, our earliest ancestors swam. Their
descendants first crawled and then walked on land. Later descendants climbed trees and, finally,
we ourselves roamed on the land again. All of these new modes of life arose as a result of new
abilities that were themselves enabled by the appearance of new genes. New genes can appear
as mutations of members of existing family members, often giving subtle variations of existing
capabilities. Entirely new capabilities can arise by a more substantial mutational variation, or by
the appearance of entirely novel gene sequences, the so-called orphan genes.
The present availability of full genome sequences of a broad range of animal species, across the
whole range of evolutionary history, enables one to identify the evolutionary stage at which
each of the eighteen thousand or so protein-coding human genes was integrated into the
developing genome. To make this identification, one looks closely at the sequence of the human
gene in which one is interested. One compares this sequence with similar gene sequences in
animals that arose earlier and earlier in evolutionary time. The age of appearance of the earliest
animal that has a gene similar in sequence, a homologue, to this human gene defines the age of
the gene in question. (We published a table of the ages of the human protein-coding genes:
T. Litman, W.D.Stein, Obtaining estimates for the ages of all the protein-coding genes and most
of the ontology-identified noncoding genes of the human genome, assigned to 19 phylostrata,
Seminars in Oncology, 46, 2019, 3-9, https://doi.org/10.1053/j.seminoncol.2018.11.002.)
With the gene ages established we can now ask, among many other questions, about the
distribution of genes across the chromosomes. Do newly recruited genes, as new animal families
emerge, distribute at random among the chromosomes or at non-random locations? To answer
this question, we extracted values for the ages of the human genes and determined their
current chromosome locations. A quantitative analysis, using the statistics of variance, showed
that the distribution of newly-added genes among, and within, the chromosomes appears to be
increasingly non-random (bunched) if one observes animals along the evolutionary series from
the precursors of the tetrapoda through to the great apes. The oldest genes are, in contrast,
randomly distributed.
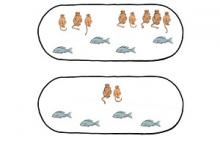
An important point needs to be noted: If one looks at the genes that first arose with the
appearance of the fishes, these genes are distributed in a highly random fashion among the
chromosomes of the fish themselves. These “fish” genes are also distributed at random among
our own chromosomes. Similarly, those that arose with the reptiles are distributed randomly
among the reptilian chromosomes, and also among our own chromosomes (but somewhat less
randomly than those that arose with the fish). In contrast, the genes that appeared with the
primates are highly bunched among the chromosomes of the early primates, and highly
bunched also in our chromosomes.
Randomization can result from chromosome evolution itself when the chromosomes join
together and then split apart during evolution, however, less and less time became available for
this process as evolution proceeds. Much of the bunching of recently-added genes arises from
the new genes being mutations of members of existing gene families, and hence located close to
genes that were recruited in the preceding time period. As examples we demonstrated the
bunching shown by the Keratin-Associated Protein gene family that determines the subtle
variation between the different forms of hair in the mammals, and we also showed bunching in
the Odour Receptor Family genes, that create the wide range of response to different smells
that is characteristic of land animals. We show that bunching can also result from the evolution
of the chromosomes themselves when, as for some members of the Keratin-associated protein
gene family, blocks of genes that had previously been on disparate chromosomes become linked
together, and their expression thus controlled in a co-ordinated way.
Read the paper – W.D. Stein, M/B/ Hoshen, During evolution from the earliest tetrapoda, newly-recruited genes are increasingly paralogues of existing genes and distribute non-randomly among the chromosomes